Overview
On this publish, we’ll overview three superior methods for bettering the efficiency and generalization energy of recurrent neural networks. By the top of the part, you’ll know most of what there may be to find out about utilizing recurrent networks with Keras. We’ll display all three ideas on a temperature-forecasting downside, the place you’ve got entry to a time sequence of information factors coming from sensors put in on the roof of a constructing, comparable to temperature, air stress, and humidity, which you employ to foretell what the temperature will likely be 24 hours after the final knowledge level. It is a pretty difficult downside that exemplifies many frequent difficulties encountered when working with time sequence.
We’ll cowl the next methods:
- Recurrent dropout — It is a particular, built-in approach to make use of dropout to battle overfitting in recurrent layers.
- Stacking recurrent layers — This will increase the representational energy of the community (at the price of larger computational hundreds).
- Bidirectional recurrent layers — These current the identical info to a recurrent community in numerous methods, growing accuracy and mitigating forgetting points.
A temperature-forecasting downside
Till now, the one sequence knowledge we’ve lined has been textual content knowledge, such because the IMDB dataset and the Reuters dataset. However sequence knowledge is discovered in lots of extra issues than simply language processing. In all of the examples on this part, you’ll play with a climate timeseries dataset recorded on the Climate Station on the Max Planck Institute for Biogeochemistry in Jena, Germany.
On this dataset, 14 completely different portions (such air temperature, atmospheric stress, humidity, wind route, and so forth) have been recorded each 10 minutes, over a number of years. The unique knowledge goes again to 2003, however this instance is restricted to knowledge from 2009–2016. This dataset is ideal for studying to work with numerical time sequence. You’ll use it to construct a mannequin that takes as enter some knowledge from the current previous (just a few days’ price of information factors) and predicts the air temperature 24 hours sooner or later.
Obtain and uncompress the info as follows:
dir.create("~/Downloads/jena_climate", recursive = TRUE)
obtain.file(
"https://s3.amazonaws.com/keras-datasets/jena_climate_2009_2016.csv.zip",
"~/Downloads/jena_climate/jena_climate_2009_2016.csv.zip"
)
unzip(
"~/Downloads/jena_climate/jena_climate_2009_2016.csv.zip",
exdir = "~/Downloads/jena_climate"
)
Let’s have a look at the info.
Observations: 420,551
Variables: 15
$ `Date Time` "01.01.2009 00:10:00", "01.01.2009 00:20:00", "...
$ `p (mbar)` 996.52, 996.57, 996.53, 996.51, 996.51, 996.50,...
$ `T (degC)` -8.02, -8.41, -8.51, -8.31, -8.27, -8.05, -7.62...
$ `Tpot (Okay)` 265.40, 265.01, 264.91, 265.12, 265.15, 265.38,...
$ `Tdew (degC)` -8.90, -9.28, -9.31, -9.07, -9.04, -8.78, -8.30...
$ `rh (%)` 93.3, 93.4, 93.9, 94.2, 94.1, 94.4, 94.8, 94.4,...
$ `VPmax (mbar)` 3.33, 3.23, 3.21, 3.26, 3.27, 3.33, 3.44, 3.44,...
$ `VPact (mbar)` 3.11, 3.02, 3.01, 3.07, 3.08, 3.14, 3.26, 3.25,...
$ `VPdef (mbar)` 0.22, 0.21, 0.20, 0.19, 0.19, 0.19, 0.18, 0.19,...
$ `sh (g/kg)` 1.94, 1.89, 1.88, 1.92, 1.92, 1.96, 2.04, 2.03,...
$ `H2OC (mmol/mol)` 3.12, 3.03, 3.02, 3.08, 3.09, 3.15, 3.27, 3.26,...
$ `rho (g/m**3)` 1307.75, 1309.80, 1310.24, 1309.19, 1309.00, 13...
$ `wv (m/s)` 1.03, 0.72, 0.19, 0.34, 0.32, 0.21, 0.18, 0.19,...
$ `max. wv (m/s)` 1.75, 1.50, 0.63, 0.50, 0.63, 0.63, 0.63, 0.50,...
$ `wd (deg)` 152.3, 136.1, 171.6, 198.0, 214.3, 192.7, 166.5...
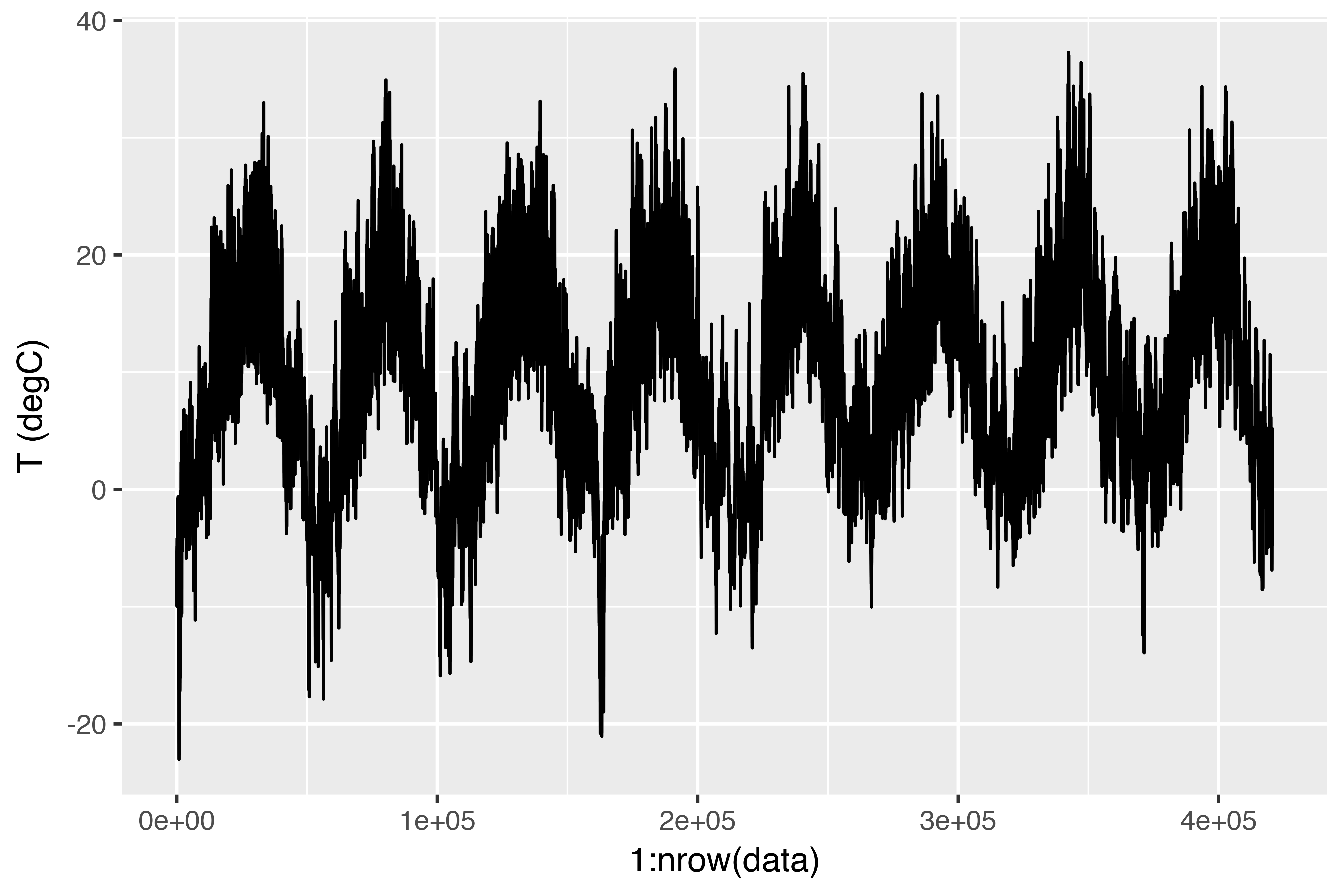
Right here is the plot of temperature (in levels Celsius) over time. On this plot, you possibly can clearly see the yearly periodicity of temperature.
Here’s a extra slim plot of the primary 10 days of temperature knowledge (see determine 6.15). As a result of the info is recorded each 10 minutes, you get 144 knowledge factors
per day.
ggplot(knowledge[1:1440,], aes(x = 1:1440, y = `T (degC)`)) + geom_line()

On this plot, you possibly can see day by day periodicity, particularly evident for the final 4 days. Additionally be aware that this 10-day interval have to be coming from a reasonably chilly winter month.
Should you have been making an attempt to foretell common temperature for the following month given just a few months of previous knowledge, the issue can be simple, as a result of dependable year-scale periodicity of the info. However wanting on the knowledge over a scale of days, the temperature seems much more chaotic. Is that this time sequence predictable at a day by day scale? Let’s discover out.
Making ready the info
The precise formulation of the issue will likely be as follows: given knowledge going way back to lookback timesteps (a timestep is 10 minutes) and sampled each steps timesteps, can you are expecting the temperature in delay timesteps? You’ll use the next parameter values:
lookback = 1440 — Observations will return 10 days.steps = 6 — Observations will likely be sampled at one knowledge level per hour.delay = 144 — Targets will likely be 24 hours sooner or later.
To get began, it’s worthwhile to do two issues:
- Preprocess the info to a format a neural community can ingest. That is simple: the info is already numerical, so that you don’t must do any vectorization. However every time sequence within the knowledge is on a distinct scale (for instance, temperature is usually between -20 and +30, however atmospheric stress, measured in mbar, is round 1,000). You’ll normalize every time sequence independently in order that all of them take small values on the same scale.
- Write a generator operate that takes the present array of float knowledge and yields batches of information from the current previous, together with a goal temperature sooner or later. As a result of the samples within the dataset are extremely redundant (pattern N and pattern N + 1 can have most of their timesteps in frequent), it could be wasteful to explicitly allocate each pattern. As an alternative, you’ll generate the samples on the fly utilizing the unique knowledge.
NOTE: Understanding generator capabilities
A generator operate is a particular sort of operate that you simply name repeatedly to acquire a sequence of values from. Usually turbines want to take care of inner state, so they’re sometimes constructed by calling one other one more operate which returns the generator operate (the setting of the operate which returns the generator is then used to trace state).
For instance, the sequence_generator() operate beneath returns a generator operate that yields an infinite sequence of numbers:
sequence_generator <- operate(begin) {
worth <- begin - 1
operate() {
worth <<- worth + 1
worth
}
}
gen <- sequence_generator(10)
gen()
[1] 10
[1] 11
The present state of the generator is the worth variable that’s outlined outdoors of the operate. Observe that superassignment (<<-) is used to replace this state from throughout the operate.
Generator capabilities can sign completion by returning the worth NULL. Nevertheless, generator capabilities handed to Keras coaching strategies (e.g. fit_generator()) ought to at all times return values infinitely (the variety of calls to the generator operate is managed by the epochs and steps_per_epoch parameters).
First, you’ll convert the R knowledge body which we learn earlier right into a matrix of floating level values (we’ll discard the primary column which included a textual content timestamp):
You’ll then preprocess the info by subtracting the imply of every time sequence and dividing by the usual deviation. You’re going to make use of the primary 200,000 timesteps as coaching knowledge, so compute the imply and normal deviation for normalization solely on this fraction of the info.
train_data <- knowledge[1:200000,]
imply <- apply(train_data, 2, imply)
std <- apply(train_data, 2, sd)
knowledge <- scale(knowledge, heart = imply, scale = std)
The code for the info generator you’ll use is beneath. It yields an inventory (samples, targets), the place samples is one batch of enter knowledge and targets is the corresponding array of goal temperatures. It takes the next arguments:
knowledge — The unique array of floating-point knowledge, which you normalized in itemizing 6.32.lookback — What number of timesteps again the enter knowledge ought to go.delay — What number of timesteps sooner or later the goal ought to be.min_index and max_index — Indices within the knowledge array that delimit which timesteps to attract from. That is helpful for conserving a phase of the info for validation and one other for testing.shuffle — Whether or not to shuffle the samples or draw them in chronological order.batch_size — The variety of samples per batch.step — The interval, in timesteps, at which you pattern knowledge. You’ll set it 6 to be able to draw one knowledge level each hour.
generator <- operate(knowledge, lookback, delay, min_index, max_index,
shuffle = FALSE, batch_size = 128, step = 6) {
if (is.null(max_index))
max_index <- nrow(knowledge) - delay - 1
i <- min_index + lookback
operate() {
if (shuffle) {
rows <- pattern(c((min_index+lookback):max_index), dimension = batch_size)
} else {
if (i + batch_size >= max_index)
i <<- min_index + lookback
rows <- c(i:min(i+batch_size-1, max_index))
i <<- i + size(rows)
}
samples <- array(0, dim = c(size(rows),
lookback / step,
dim(knowledge)[[-1]]))
targets <- array(0, dim = c(size(rows)))
for (j in 1:size(rows)) {
indices <- seq(rows[[j]] - lookback, rows[[j]]-1,
size.out = dim(samples)[[2]])
samples[j,,] <- knowledge[indices,]
targets[[j]] <- knowledge[rows[[j]] + delay,2]
}
checklist(samples, targets)
}
}
The i variable incorporates the state that tracks subsequent window of information to return, so it’s up to date utilizing superassignment (e.g. i <<- i + size(rows)).
Now, let’s use the summary generator operate to instantiate three turbines: one for coaching, one for validation, and one for testing. Every will have a look at completely different temporal segments of the unique knowledge: the coaching generator seems on the first 200,000 timesteps, the validation generator seems on the following 100,000, and the check generator seems on the the rest.
lookback <- 1440
step <- 6
delay <- 144
batch_size <- 128
train_gen <- generator(
knowledge,
lookback = lookback,
delay = delay,
min_index = 1,
max_index = 200000,
shuffle = TRUE,
step = step,
batch_size = batch_size
)
val_gen = generator(
knowledge,
lookback = lookback,
delay = delay,
min_index = 200001,
max_index = 300000,
step = step,
batch_size = batch_size
)
test_gen <- generator(
knowledge,
lookback = lookback,
delay = delay,
min_index = 300001,
max_index = NULL,
step = step,
batch_size = batch_size
)
# What number of steps to attract from val_gen to be able to see your entire validation set
val_steps <- (300000 - 200001 - lookback) / batch_size
# What number of steps to attract from test_gen to be able to see your entire check set
test_steps <- (nrow(knowledge) - 300001 - lookback) / batch_size
A typical-sense, non-machine-learning baseline
Earlier than you begin utilizing black-box deep-learning fashions to resolve the temperature-prediction downside, let’s attempt a easy, commonsense strategy. It’ll function a sanity examine, and it’ll set up a baseline that you simply’ll need to beat to be able to display the usefulness of more-advanced machine-learning fashions. Such commonsense baselines may be helpful whenever you’re approaching a brand new downside for which there isn’t any recognized answer (but). A traditional instance is that of unbalanced classification duties, the place some lessons are rather more frequent than others. In case your dataset incorporates 90% cases of sophistication A and 10% cases of sophistication B, then a commonsense strategy to the classification job is to at all times predict “A” when introduced with a brand new pattern. Such a classifier is 90% correct total, and any learning-based strategy ought to due to this fact beat this 90% rating to be able to display usefulness. Typically, such elementary baselines can show surprisingly arduous to beat.
On this case, the temperature time sequence can safely be assumed to be steady (the temperatures tomorrow are more likely to be near the temperatures as we speak) in addition to periodical with a day by day interval. Thus a commonsense strategy is to at all times predict that the temperature 24 hours from now will likely be equal to the temperature proper now. Let’s consider this strategy, utilizing the imply absolute error (MAE) metric:
Right here’s the analysis loop.
library(keras)
evaluate_naive_method <- operate() {
batch_maes <- c()
for (step in 1:val_steps) {
c(samples, targets) %<-% val_gen()
preds <- samples[,dim(samples)[[2]],2]
mae <- imply(abs(preds - targets))
batch_maes <- c(batch_maes, mae)
}
print(imply(batch_maes))
}
evaluate_naive_method()
This yields an MAE of 0.29. As a result of the temperature knowledge has been normalized to be centered on 0 and have a typical deviation of 1, this quantity isn’t instantly interpretable. It interprets to a median absolute error of 0.29 x temperature_std levels Celsius: 2.57˚C.
celsius_mae <- 0.29 * std[[2]]
That’s a reasonably large common absolute error. Now the sport is to make use of your information of deep studying to do higher.
A primary machine-learning strategy
In the identical approach that it’s helpful to determine a commonsense baseline earlier than making an attempt machine-learning approaches, it’s helpful to attempt easy, low-cost machine-learning fashions (comparable to small, densely linked networks) earlier than wanting into sophisticated and computationally costly fashions comparable to RNNs. That is the easiest way to verify any additional complexity you throw on the downside is respectable and delivers actual advantages.
The next itemizing reveals a completely linked mannequin that begins by flattening the info after which runs it by two dense layers. Observe the shortage of activation operate on the final dense layer, which is typical for a regression downside. You utilize MAE because the loss. Since you consider on the very same knowledge and with the very same metric you probably did with the common sense strategy, the outcomes will likely be immediately comparable.
library(keras)
mannequin <- keras_model_sequential() %>%
layer_flatten(input_shape = c(lookback / step, dim(knowledge)[-1])) %>%
layer_dense(items = 32, activation = "relu") %>%
layer_dense(items = 1)
mannequin %>% compile(
optimizer = optimizer_rmsprop(),
loss = "mae"
)
historical past <- mannequin %>% fit_generator(
train_gen,
steps_per_epoch = 500,
epochs = 20,
validation_data = val_gen,
validation_steps = val_steps
)
Let’s show the loss curves for validation and coaching.

A few of the validation losses are near the no-learning baseline, however not reliably. This goes to indicate the benefit of getting this baseline within the first place: it seems to be not simple to outperform. Your frequent sense incorporates loads of priceless info {that a} machine-learning mannequin doesn’t have entry to.
You might surprise, if a easy, well-performing mannequin exists to go from the info to the targets (the common sense baseline), why doesn’t the mannequin you’re coaching discover it and enhance on it? As a result of this easy answer isn’t what your coaching setup is on the lookout for. The area of fashions wherein you’re trying to find an answer – that’s, your speculation area – is the area of all doable two-layer networks with the configuration you outlined. These networks are already pretty sophisticated. While you’re on the lookout for an answer with an area of sophisticated fashions, the straightforward, well-performing baseline could also be unlearnable, even when it’s technically a part of the speculation area. That may be a fairly vital limitation of machine studying on the whole: until the training algorithm is hardcoded to search for a particular form of easy mannequin, parameter studying can typically fail to discover a easy answer to a easy downside.
A primary recurrent baseline
The primary absolutely linked strategy didn’t do nicely, however that doesn’t imply machine studying isn’t relevant to this downside. The earlier strategy first flattened the time sequence, which eliminated the notion of time from the enter knowledge. Let’s as an alternative have a look at the info as what it’s: a sequence, the place causality and order matter. You’ll attempt a recurrent-sequence processing mannequin – it ought to be the proper match for such sequence knowledge, exactly as a result of it exploits the temporal ordering of information factors, in contrast to the primary strategy.
As an alternative of the LSTM layer launched within the earlier part, you’ll use the GRU layer, developed by Chung et al. in 2014. Gated recurrent unit (GRU) layers work utilizing the identical precept as LSTM, however they’re considerably streamlined and thus cheaper to run (though they might not have as a lot representational energy as LSTM). This trade-off between computational expensiveness and representational energy is seen all over the place in machine studying.
mannequin <- keras_model_sequential() %>%
layer_gru(items = 32, input_shape = checklist(NULL, dim(knowledge)[[-1]])) %>%
layer_dense(items = 1)
mannequin %>% compile(
optimizer = optimizer_rmsprop(),
loss = "mae"
)
historical past <- mannequin %>% fit_generator(
train_gen,
steps_per_epoch = 500,
epochs = 20,
validation_data = val_gen,
validation_steps = val_steps
)
The outcomes are plotted beneath. Significantly better! You may considerably beat the common sense baseline, demonstrating the worth of machine studying in addition to the prevalence of recurrent networks in comparison with sequence-flattening dense networks on this sort of job.

The brand new validation MAE of ~0.265 (earlier than you begin considerably overfitting) interprets to a imply absolute error of two.35˚C after denormalization. That’s a stable acquire on the preliminary error of two.57˚C, however you in all probability nonetheless have a little bit of a margin for enchancment.
Utilizing recurrent dropout to battle overfitting
It’s evident from the coaching and validation curves that the mannequin is overfitting: the coaching and validation losses begin to diverge significantly after just a few epochs. You’re already aware of a traditional method for preventing this phenomenon: dropout, which randomly zeros out enter items of a layer to be able to break happenstance correlations within the coaching knowledge that the layer is uncovered to. However tips on how to appropriately apply dropout in recurrent networks isn’t a trivial query. It has lengthy been recognized that making use of dropout earlier than a recurrent layer hinders studying relatively than serving to with regularization. In 2015, Yarin Gal, as a part of his PhD thesis on Bayesian deep studying, decided the correct approach to make use of dropout with a recurrent community: the identical dropout masks (the identical sample of dropped items) ought to be utilized at each timestep, as an alternative of a dropout masks that varies randomly from timestep to timestep. What’s extra, to be able to regularize the representations shaped by the recurrent gates of layers comparable to layer_gru and layer_lstm, a temporally fixed dropout masks ought to be utilized to the inside recurrent activations of the layer (a recurrent dropout masks). Utilizing the identical dropout masks at each timestep permits the community to correctly propagate its studying error by time; a temporally random dropout masks would disrupt this error sign and be dangerous to the training course of.
Yarin Gal did his analysis utilizing Keras and helped construct this mechanism immediately into Keras recurrent layers. Each recurrent layer in Keras has two dropout-related arguments: dropout, a float specifying the dropout charge for enter items of the layer, and recurrent_dropout, specifying the dropout charge of the recurrent items. Let’s add dropout and recurrent dropout to the layer_gru and see how doing so impacts overfitting. As a result of networks being regularized with dropout at all times take longer to totally converge, you’ll practice the community for twice as many epochs.
mannequin <- keras_model_sequential() %>%
layer_gru(items = 32, dropout = 0.2, recurrent_dropout = 0.2,
input_shape = checklist(NULL, dim(knowledge)[[-1]])) %>%
layer_dense(items = 1)
mannequin %>% compile(
optimizer = optimizer_rmsprop(),
loss = "mae"
)
historical past <- mannequin %>% fit_generator(
train_gen,
steps_per_epoch = 500,
epochs = 40,
validation_data = val_gen,
validation_steps = val_steps
)
The plot beneath reveals the outcomes. Success! You’re not overfitting throughout the first 20 epochs. However though you’ve got extra secure analysis scores, your greatest scores aren’t a lot decrease than they have been beforehand.

Stacking recurrent layers
Since you’re not overfitting however appear to have hit a efficiency bottleneck, you need to take into account growing the capability of the community. Recall the outline of the common machine-learning workflow: it’s usually a good suggestion to extend the capability of your community till overfitting turns into the first impediment (assuming you’re already taking primary steps to mitigate overfitting, comparable to utilizing dropout). So long as you aren’t overfitting too badly, you’re probably below capability.
Rising community capability is usually achieved by growing the variety of items within the layers or including extra layers. Recurrent layer stacking is a traditional approach to construct more-powerful recurrent networks: for example, what presently powers the Google Translate algorithm is a stack of seven giant LSTM layers – that’s large.
To stack recurrent layers on prime of one another in Keras, all intermediate layers ought to return their full sequence of outputs (a 3D tensor) relatively than their output on the final timestep. That is achieved by specifying return_sequences = TRUE.
mannequin <- keras_model_sequential() %>%
layer_gru(items = 32,
dropout = 0.1,
recurrent_dropout = 0.5,
return_sequences = TRUE,
input_shape = checklist(NULL, dim(knowledge)[[-1]])) %>%
layer_gru(items = 64, activation = "relu",
dropout = 0.1,
recurrent_dropout = 0.5) %>%
layer_dense(items = 1)
mannequin %>% compile(
optimizer = optimizer_rmsprop(),
loss = "mae"
)
historical past <- mannequin %>% fit_generator(
train_gen,
steps_per_epoch = 500,
epochs = 40,
validation_data = val_gen,
validation_steps = val_steps
)
The determine beneath reveals the outcomes. You may see that the added layer does enhance the outcomes a bit, although not considerably. You may draw two conclusions:
- Since you’re nonetheless not overfitting too badly, you might safely improve the scale of your layers in a quest for validation-loss enchancment. This has a non-negligible computational price, although.
- Including a layer didn’t assist by a major issue, so it’s possible you’ll be seeing diminishing returns from growing community capability at this level.

Utilizing bidirectional RNNs
The final method launched on this part is named bidirectional RNNs. A bidirectional RNN is a typical RNN variant that may provide better efficiency than a daily RNN on sure duties. It’s often utilized in natural-language processing – you might name it the Swiss Military knife of deep studying for natural-language processing.
RNNs are notably order dependent, or time dependent: they course of the timesteps of their enter sequences so as, and shuffling or reversing the timesteps can utterly change the representations the RNN extracts from the sequence. That is exactly the rationale they carry out nicely on issues the place order is significant, such because the temperature-forecasting downside. A bidirectional RNN exploits the order sensitivity of RNNs: it consists of utilizing two common RNNs, such because the layer_gru and layer_lstm you’re already aware of, every of which processes the enter sequence in a single route (chronologically and antichronologically), after which merging their representations. By processing a sequence each methods, a bidirectional RNN can catch patterns that could be neglected by a unidirectional RNN.
Remarkably, the truth that the RNN layers on this part have processed sequences in chronological order (older timesteps first) could have been an arbitrary resolution. No less than, it’s a choice we made no try to query thus far. Might the RNNs have carried out nicely sufficient in the event that they processed enter sequences in antichronological order, for example (newer timesteps first)? Let’s do that in observe and see what occurs. All it’s worthwhile to do is write a variant of the info generator the place the enter sequences are reverted alongside the time dimension (exchange the final line with checklist(samples[,ncol(samples):1,], targets)). Coaching the identical one-GRU-layer community that you simply used within the first experiment on this part, you get the outcomes proven beneath.

The reversed-order GRU underperforms even the common sense baseline, indicating that on this case, chronological processing is necessary to the success of your strategy. This makes excellent sense: the underlying GRU layer will sometimes be higher at remembering the current previous than the distant previous, and naturally the newer climate knowledge factors are extra predictive than older knowledge factors for the issue (that’s what makes the common sense baseline pretty robust). Thus the chronological model of the layer is sure to outperform the reversed-order model. Importantly, this isn’t true for a lot of different issues, together with pure language: intuitively, the significance of a phrase in understanding a sentence isn’t often depending on its place within the sentence. Let’s attempt the identical trick on the LSTM IMDB instance from part 6.2.
library(keras)
# Variety of phrases to think about as options
max_features <- 10000
# Cuts off texts after this variety of phrases
maxlen <- 500
imdb <- dataset_imdb(num_words = max_features)
c(c(x_train, y_train), c(x_test, y_test)) %<-% imdb
# Reverses sequences
x_train <- lapply(x_train, rev)
x_test <- lapply(x_test, rev)
# Pads sequences
x_train <- pad_sequences(x_train, maxlen = maxlen) <4>
x_test <- pad_sequences(x_test, maxlen = maxlen)
mannequin <- keras_model_sequential() %>%
layer_embedding(input_dim = max_features, output_dim = 128) %>%
layer_lstm(items = 32) %>%
layer_dense(items = 1, activation = "sigmoid")
mannequin %>% compile(
optimizer = "rmsprop",
loss = "binary_crossentropy",
metrics = c("acc")
)
historical past <- mannequin %>% match(
x_train, y_train,
epochs = 10,
batch_size = 128,
validation_split = 0.2
)
You get efficiency almost an identical to that of the chronological-order LSTM. Remarkably, on such a textual content dataset, reversed-order processing works simply in addition to chronological processing, confirming the
speculation that, though phrase order does matter in understanding language, which order you employ isn’t essential. Importantly, an RNN educated on reversed sequences will be taught completely different representations than one educated on the unique sequences, a lot as you’ll have completely different psychological fashions if time flowed backward in the true world – in case you lived a life the place you died in your first day and have been born in your final day. In machine studying, representations which can be completely different but helpful are at all times price exploiting, and the extra they differ, the higher: they provide a unique approach from which to have a look at your knowledge, capturing points of the info that have been missed by different approaches, and thus they will help enhance efficiency on a job. That is the instinct behind ensembling, an idea we’ll discover in chapter 7.
A bidirectional RNN exploits this concept to enhance on the efficiency of chronological-order RNNs. It seems at its enter sequence each methods, acquiring doubtlessly richer representations and capturing patterns that will have been missed by the chronological-order model alone.

To instantiate a bidirectional RNN in Keras, you employ the bidirectional() operate, which takes a recurrent layer occasion as an argument. The bidirectional() operate creates a second, separate occasion of this recurrent layer and makes use of one occasion for processing the enter sequences in chronological order and the opposite occasion for processing the enter sequences in reversed order. Let’s attempt it on the IMDB sentiment-analysis job.
mannequin <- keras_model_sequential() %>%
layer_embedding(input_dim = max_features, output_dim = 32) %>%
bidirectional(
layer_lstm(items = 32)
) %>%
layer_dense(items = 1, activation = "sigmoid")
mannequin %>% compile(
optimizer = "rmsprop",
loss = "binary_crossentropy",
metrics = c("acc")
)
historical past <- mannequin %>% match(
x_train, y_train,
epochs = 10,
batch_size = 128,
validation_split = 0.2
)
It performs barely higher than the common LSTM you tried within the earlier part, attaining over 89% validation accuracy. It additionally appears to overfit extra shortly, which is unsurprising as a result of a bidirectional layer has twice as many parameters as a chronological LSTM. With some regularization, the bidirectional strategy would probably be a powerful performer on this job.
Now let’s attempt the identical strategy on the temperature prediction job.
mannequin <- keras_model_sequential() %>%
bidirectional(
layer_gru(items = 32), input_shape = checklist(NULL, dim(knowledge)[[-1]])
) %>%
layer_dense(items = 1)
mannequin %>% compile(
optimizer = optimizer_rmsprop(),
loss = "mae"
)
historical past <- mannequin %>% fit_generator(
train_gen,
steps_per_epoch = 500,
epochs = 40,
validation_data = val_gen,
validation_steps = val_steps
)
This performs about in addition to the common layer_gru. It’s simple to know why: all of the predictive capability should come from the chronological half of the community, as a result of the antichronological half is understood to be severely underperforming on this job (once more, as a result of the current previous issues rather more than the distant previous on this case).
Going even additional
There are various different issues you might attempt, to be able to enhance efficiency on the temperature-forecasting downside:
- Regulate the variety of items in every recurrent layer within the stacked setup. The present decisions are largely arbitrary and thus in all probability suboptimal.
- Regulate the training charge utilized by the
RMSprop optimizer.
- Strive utilizing
layer_lstm as an alternative of layer_gru.
- Strive utilizing an even bigger densely linked regressor on prime of the recurrent layers: that’s, an even bigger dense layer or perhaps a stack of dense layers.
- Don’t neglect to finally run the best-performing fashions (when it comes to validation MAE) on the check set! In any other case, you’ll develop architectures which can be overfitting to the validation set.
As at all times, deep studying is extra an artwork than a science. We are able to present pointers that recommend what’s more likely to work or not work on a given downside, however, in the end, each downside is exclusive; you’ll have to judge completely different methods empirically. There’s presently no concept that may inform you upfront exactly what you need to do to optimally resolve an issue. It’s essential to iterate.
Wrapping up
Right here’s what you need to take away from this part:
- As you first discovered in chapter 4, when approaching a brand new downside, it’s good to first set up commonsense baselines in your metric of selection. Should you don’t have a baseline to beat, you possibly can’t inform whether or not you’re making actual progress.
- Strive easy fashions earlier than costly ones, to justify the extra expense. Typically a easy mannequin will grow to be your only option.
- When you’ve got knowledge the place temporal ordering issues, recurrent networks are an excellent match and simply outperform fashions that first flatten the temporal knowledge.
- To make use of dropout with recurrent networks, you need to use a time-constant dropout masks and recurrent dropout masks. These are constructed into Keras recurrent layers, so all you need to do is use the
dropout and recurrent_dropout arguments of recurrent layers.
- Stacked RNNs present extra representational energy than a single RNN layer. They’re additionally rather more costly and thus not at all times price it. Though they provide clear good points on complicated issues (comparable to machine translation), they might not at all times be related to smaller, less complicated issues.
- Bidirectional RNNs, which have a look at a sequence each methods, are helpful on natural-language processing issues. However they aren’t robust performers on sequence knowledge the place the current previous is rather more informative than the start of the sequence.
NOTE: Markets and machine studying
Some readers are sure to need to take the methods we’ve launched right here and take a look at them on the issue of forecasting the longer term worth of securities on the inventory market (or foreign money trade charges, and so forth). Markets have very completely different statistical traits than pure phenomena comparable to climate patterns. Making an attempt to make use of machine studying to beat markets, whenever you solely have entry to publicly obtainable knowledge, is a troublesome endeavor, and also you’re more likely to waste your time and assets with nothing to indicate for it.
At all times do not forget that in the case of markets, previous efficiency is not predictor of future returns – wanting within the rear-view mirror is a nasty approach to drive. Machine studying, alternatively, is relevant to datasets the place the previous is predictor of the longer term.
Take pleasure in this weblog? Get notified of recent posts by electronic mail:
Posts additionally obtainable at r-bloggers