In my final put up, we realized easy methods to import the uncooked COVID-19 knowledge from the Johns Hopkins GitHub repository. This put up will exhibit easy methods to convert the uncooked knowledge to time-series knowledge. We’ll additionally create some tables and graphs alongside the way in which.
Let’s have a look at the uncooked COVID-19 knowledge that we saved earlier.
. use covid19_raw, clear
. describe
Comprises knowledge from covid19_raw.dta
obs: 11,341
vars: 12 24 Mar 2020 12:55
------------------------------------------------------------------------
storage show worth
variable title sort format label variable label
------------------------------------------------------------------------
provincestate str43 %43s Province/State
countryregion str32 %32s Nation/Area
lastupdate str19 %19s Final Replace
confirmed lengthy %8.0g Confirmed
deaths int %8.0g Deaths
recovered lengthy %8.0g Recovered
latitude float %9.0g Latitude
longitude float %9.0g Longitude
fips lengthy %12.0g FIPS
admin2 str21 %21s Admin2
lively lengthy %12.0g Lively
combined_key str44 %44s Combined_Key
------------------------------------------------------------------------
Sorted by:
Our dataset incorporates 11,341 observations on 12 variables. Let’s checklist the primary 5 observations for lastupdate.
. checklist lastupdate in 1/5
+-----------------+
| lastupdate |
|-----------------|
1. | 1/22/2020 17:00 |
2. | 1/22/2020 17:00 |
3. | 1/22/2020 17:00 |
4. | 1/22/2020 17:00 |
5. | 1/22/2020 17:00 |
+-----------------+
lastupdate is the replace time and date for every commentary within the dataset. The information embrace the date adopted by an area adopted by time.
Let’s additionally have a look at the final 5 observations within the dataset.
. checklist lastupdate in -5/l
+---------------------+
| lastupdate |
|---------------------|
11337. | 2020-03-23 23:19:21 |
11338. | 2020-03-23 23:19:21 |
11339. | 2020-03-23 23:19:21 |
11340. | 2020-03-23 23:19:21 |
11341. | 2020-03-23 23:19:21 |
+---------------------+
The final 5 observations comprise related info however in a unique format. Sadly, the info for lastupdate weren’t saved persistently within the uncooked knowledge information. We may look at the other ways dates have been saved in lastupdate throughout the totally different information and develop a method to extract the dates. However we all know the date for every file as a result of it’s a part of the title for every uncooked knowledge file. In my earlier posts, we created a persistently formatted date once we imported the uncooked knowledge. A easy resolution could be to avoid wasting that date as a variable once we import the uncooked knowledge information. Let’s generate a variable named tempdate in every uncooked knowledge file.
native URL = "https://uncooked.githubusercontent.com/CSSEGISandData/COVID-19/grasp/csse_covid_19_data/csse_covid_19_daily_reports/"
forvalues month = 1/12 {
forvalues day = 1/31 {
native month = string(`month', "%02.0f")
native day = string(`day', "%02.0f")
native 12 months = "2020"
native right this moment = "`month'-`day'-`12 months'"
native FileName = "`URL'`right this moment'.csv"
clear
seize import delimited "`FileName'"
seize affirm variable ïprovincestate
if _rc == 0 {
rename ïprovincestate provincestate
label variable provincestate "Province/State"
}
seize rename province_state provincestate
seize rename country_region countryregion
seize rename last_update lastupdate
seize rename lat latitude
seize rename lengthy longitude
generate tempdate = "`right this moment'"
seize save "`right this moment'", change
}
}
clear
forvalues month = 1/12 {
forvalues day = 1/31 {
native month = string(`month', "%02.0f")
native day = string(`day', "%02.0f")
native 12 months = "2020"
native right this moment = "`month'-`day'-`12 months'"
seize append utilizing "`right this moment'"
}
}
We will verify our work by checklisting the observations. Under, I’ve checklisted the primary 5 observations and the final 5 observations in our mixed uncooked dataset.
. checklist tempdate in 1/5
+------------+
| tempdate |
|------------|
1. | 01-22-2020 |
2. | 01-22-2020 |
3. | 01-22-2020 |
4. | 01-22-2020 |
5. | 01-22-2020 |
+------------+
. checklist tempdate in -5/l
+------------+
| tempdate |
|------------|
11337. | 03-23-2020 |
11338. | 03-23-2020 |
11339. | 03-23-2020 |
11340. | 03-23-2020 |
11341. | 03-23-2020 |
+------------+
The date knowledge saved in tempdate are saved persistently, however the knowledge are nonetheless saved as a string. We will use the date() operate to transform tempdate to a quantity. The date(s1,s2) operate returns a quantity primarily based on two arguments, s1 and s2. The argument s1 is the string we want to act upon and the argument s2 is the order of the day, month, and 12 months in s1. Our tempdate variable is saved with the month first, the day second, and the 12 months third. So we are able to sort s2 as MDY, which signifies that Month is adopted by Day, which is adopted by Year. We will use the date() operate beneath to transform the string date 03-23-2020 to a quantity.
. show date("03-23-2020", "MDY")
21997
The date() operate returned the quantity 21997. That doesn’t appear to be a date to you and me, nevertheless it signifies the variety of days since January 1, 1960. The instance beneath exhibits that 01-01-1960 is the 0 for our time knowledge.
. show date("01-01-1960", "MDY")
0
We will change the way in which the quantity is displayed by making use of a date format to the quantity.
. show %tdNN/DD/CCYY date("03-23-2020", "MDY")
03/23/2020
Let’s use date() to generate a brand new variable named date.
. generate date = date(tempdate, "MDY")
. checklist lastupdate tempdate date in -5/l
+------------------------------------------+
| lastupdate tempdate date |
|------------------------------------------|
11337. | 2020-03-23 23:19:21 03-23-2020 21997 |
11338. | 2020-03-23 23:19:21 03-23-2020 21997 |
11339. | 2020-03-23 23:19:21 03-23-2020 21997 |
11340. | 2020-03-23 23:19:21 03-23-2020 21997 |
11341. | 2020-03-23 23:19:21 03-23-2020 21997 |
+------------------------------------------+
Subsequent, we are able to use format to show the numbers in date in a manner that appears acquainted to you and me.
. format date %tdNN/DD/CCYY
. checklist lastupdate tempdate date in -5/l
+-----------------------------------------------+
| lastupdate tempdate date |
|-----------------------------------------------|
11337. | 2020-03-23 23:19:21 03-23-2020 03/23/2020 |
11338. | 2020-03-23 23:19:21 03-23-2020 03/23/2020 |
11339. | 2020-03-23 23:19:21 03-23-2020 03/23/2020 |
11340. | 2020-03-23 23:19:21 03-23-2020 03/23/2020 |
11341. | 2020-03-23 23:19:21 03-23-2020 03/23/2020 |
+-----------------------------------------------+
Now, we now have a date variable in our dataset that can be utilized with Stata’s time-series options and for different calculations.
Let’s save this dataset in order that we don’t must obtain the uncooked knowledge for every of the next examples.
. save covid19_date, change file covid19_date.dta saved
Create time-series knowledge
Let’s hold the info for the USA and checklist the info for January 26, 2020.
. hold if countryregion=="US"
(6,545 observations deleted)
. checklist date confirmed deaths recovered ///
if date==date("1/26/2020", "MDY"), abbreviate(13)
+---------------------------------------------+
| date confirmed deaths recovered |
|---------------------------------------------|
7. | 01/26/2020 1 . . |
8. | 01/26/2020 1 . . |
9. | 01/26/2020 2 . . |
10. | 01/26/2020 1 . . |
+---------------------------------------------+
There have been 4 observations for the USA on January 26, 2020, and 5 confirmed circumstances of COVID-19. We want our time-series knowledge to comprise all 5 confirmed circumstances in a single commentary for a single date. We will use collapse to mixture the info by date.
. collapse (sum) confirmed deaths recovered, by(date)
. checklist date confirmed deaths recovered ///
if date==date("1/26/2020", "MDY"), abbreviate(13)
+---------------------------------------------+
| date confirmed deaths recovered |
|---------------------------------------------|
5. | 01/26/2020 5 0 0 |
+---------------------------------------------+
Let’s use the info for all nations and collapse the dataset by date.
. use covid19_date, clear . collapse (sum) confirmed deaths recovered, by(date)
We will describe our new dataset and see that it incorporates the variables date, confirmed, deaths, and recovered. The variable labels inform us that confirmed, deaths, and recovered are the sum of every variable for every worth of date.
. describe
Comprises knowledge
obs: 60
vars: 4
------------------------------------------------------------------------
storage show worth
variable title sort format label variable label
------------------------------------------------------------------------
date float %td..
confirmed double %8.0g (sum) confirmed
deaths double %8.0g (sum) deaths
recovered double %8.0g (sum) recovered
------------------------------------------------------------------------
Sorted by: date
Notice: Dataset has modified since final saved.
We will checklist the primary 5 observations to confirm that we now have one worth of every variable for every date.
. checklist in 1/5, abbreviate(9)
+---------------------------------------------+
| date confirmed deaths recovered |
|---------------------------------------------|
1. | 01/22/2020 555 17 28 |
2. | 01/23/2020 653 18 30 |
3. | 01/24/2020 941 26 36 |
4. | 01/25/2020 1438 42 39 |
5. | 01/26/2020 2118 56 52 |
+---------------------------------------------+
The counts are massive and can possible develop bigger. We will make our knowledge simpler to learn by utilizing format so as to add commas.
. format %8.0fc confirmed deaths recovered
. checklist, abbreviate(9)
+---------------------------------------------+
| date confirmed deaths recovered |
|---------------------------------------------|
1. | 01/22/2020 555 17 28 |
2. | 01/23/2020 653 18 30 |
3. | 01/24/2020 941 26 36 |
4. | 01/25/2020 1,438 42 39 |
5. | 01/26/2020 2,118 56 52 |
|---------------------------------------------|
6. | 01/27/2020 2,927 82 61 |
(Output omitted)
58. | 03/19/2020 242,713 9,867 84,962 |
59. | 03/20/2020 272,167 11,299 87,403 |
60. | 03/21/2020 304,528 12,973 91,676 |
|---------------------------------------------|
61. | 03/22/2020 335,957 14,634 97,882 |
62. | 03/23/2020 378,287 16,497 100,958 |
+---------------------------------------------+
Now, we are able to use tsset to specify the construction of our time-series knowledge, which is able to enable us to make use of Stata’s time-series options.
. tsset date, day by day
time variable: date, 01/22/2020 to 03/23/2020
delta: 1 day
Subsequent, I wish to calculate the variety of new circumstances reported every day. That is straightforward utilizing time-series operators. The time-series operator D.varname calculates the distinction between an commentary and the previous commentary in varname. Let’s contemplate the worth of confirmed for commentary 2, which is 653. The previous worth of confirmed, commentary 1, is 555. So D.confirmed for commentary 2 is 653 – 555, which equals 98. The information for newcases in commentary 1 are lacking as a result of there aren’t any knowledge previous commentary 1.
. generate newcases = D.confirmed
(1 lacking worth generated)
. checklist, abbreviate(9)
+--------------------------------------------------------+
| date confirmed deaths recovered newcases |
|--------------------------------------------------------|
1. | 01/22/2020 555 17 28 . |
2. | 01/23/2020 653 18 30 98 |
3. | 01/24/2020 941 26 36 288 |
4. | 01/25/2020 1,438 42 39 497 |
5. | 01/26/2020 2,118 56 52 680 |
|--------------------------------------------------------|
6. | 01/27/2020 2,927 82 61 809 |
(Output omitted)
58. | 03/19/2020 242,713 9,867 84,962 27798 |
59. | 03/20/2020 272,167 11,299 87,403 29454 |
60. | 03/21/2020 304,528 12,973 91,676 32361 |
|--------------------------------------------------------|
61. | 03/22/2020 335,957 14,634 97,882 31429 |
62. | 03/23/2020 378,287 16,497 100,958 42330 |
+--------------------------------------------------------+
Let’s create a time-series plot for the variety of confirmed circumstances for all nations mixed.
tsline confirmed, title(World Confirmed COVID-19 Circumstances)
Determine 1: World confirmed COVID-19 circumstances
Create time-series knowledge for a number of nations
In some unspecified time in the future we could want to examine the info for various nations. There are a number of methods we may do that. We may create a separate dataset for every nation and merge the datasets. We may additionally create a number of knowledge frames and use frlink to hyperlink the frames. I’m going to point out you the way to do that utilizing collapse and reshape. Let’s start by opening our uncooked time knowledge and tabulating countryregion. There are 210 nations within the desk, so I’ve eliminated many rows to shorten the desk.
. use covid19_date, clear
. tab countryregion
Nation/Area | Freq. % Cum.
---------------------------------+-----------------------------------
Azerbaijan | 1 0.01 0.01
Afghanistan | 29 0.26 0.26
Albania | 15 0.13 0.40
(rows omitted)
China | 429 3.78 14.29
(rows omitted)
Italy | 53 0.47 24.62
(rows omitted)
Mainland China | 1,517 13.38 41.24
(rows omitted)
US | 4,796 42.29 97.78
(rows omitted)
Viet Nam | 1 0.01 99.32
Vietnam | 60 0.53 99.85
Zambia | 6 0.05 99.90
Zimbabwe | 4 0.04 99.94
occupied Palestinian territory | 7 0.06 100.00
---------------------------------+-----------------------------------
Whole | 11,341 100.00
Two classes embrace China: “China” and “Mainland China”. Nearer inspection of the uncooked knowledge exhibits that the title was modified from “Mainland China” to “China” after March 12, 2020. I’m going to mix the info by renaming the class “Mainland China”.
. change countryregion = "China" if countryregion=="Mainland China" (1,517 actual adjustments made)
Subsequent, I’m going to hold the observations from China, Italy, and the USA utilizing inlist().
. hold if inlist(countryregion, "China", "US", "Italy")
(4,546 observations deleted)
. tab countryregion
Nation/Area | Freq. % Cum.
---------------------------------+-----------------------------------
China | 1,946 28.64 28.64
Italy | 53 0.78 29.42
US | 4,796 70.58 100.00
---------------------------------+-----------------------------------
Whole | 6,795 100.00
Now, we are able to collapse the info by each date and countryregion.
. collapse (sum) confirmed deaths recovered, by(date countryregion)
. checklist date countryregion confirmed deaths recovered ///
in -9/l, sepby(date) abbreviate(13)
+-------------------------------------------------------------+
| date countryregion confirmed deaths recovered |
|-------------------------------------------------------------|
169. | 03/21/2020 China 81305 3259 71857 |
170. | 03/21/2020 Italy 53578 4825 6072 |
171. | 03/21/2020 US 25493 307 171 |
|-------------------------------------------------------------|
172. | 03/22/2020 China 81397 3265 72362 |
173. | 03/22/2020 Italy 59138 5476 7024 |
174. | 03/22/2020 US 33276 417 178 |
|-------------------------------------------------------------|
175. | 03/23/2020 China 81496 3274 72819 |
176. | 03/23/2020 Italy 63927 6077 7432 |
177. | 03/23/2020 US 43667 552 0 |
+-------------------------------------------------------------+
Our new dataset incorporates one commentary for every date for every nation. The variable countryregion is saved as a string variable, and I occur to know that we are going to want a numeric variable for a few of the instructions we are going to use shortly. I’ll omit the main points of my trials and errors and easily present you easy methods to use encode to create a labeled, numeric variable named nation.
. encode countryregion, gen(nation)
. checklist date countryregion nation ///
in -9/l, sepby(date) abbreviate(13)
+--------------------------------------+
| date countryregion nation |
|--------------------------------------|
169. | 03/21/2020 China China |
170. | 03/21/2020 Italy Italy |
171. | 03/21/2020 US US |
|--------------------------------------|
172. | 03/22/2020 China China |
173. | 03/22/2020 Italy Italy |
174. | 03/22/2020 US US |
|--------------------------------------|
175. | 03/23/2020 China China |
176. | 03/23/2020 Italy Italy |
177. | 03/23/2020 US US |
+--------------------------------------+
The variables countryregion and nation look the identical once we checklist the info. However nation is a numeric variable with worth labels that have been created by encode. You possibly can sort label checklist to view the classes of nation.
. label checklist nation
nation:
1 China
2 Italy
3 US
Our knowledge are in lengthy type as a result of the time sequence for the three nations are stacked on high of one another. We may use tsset to inform Stata that we now have time-series knowledge with panels (nations).
. tsset nation date, day by day
panel variable: nation (unbalanced)
time variable: date, 01/22/2020 to 03/23/2020
delta: 1 day
You might want to save this model of the dataset should you plan to make use of Stata’s options for time-series evaluation with panel knowledge.
. save covide19_long file covid19_long.dta saved
Use reshape to create huge time-series knowledge for a number of nations
You would possibly choose to have your knowledge in huge format in order that the info for every nation are facet by facet. We will use reshape to do that. Let’s hold solely the info we are going to use earlier than we use reshape.
. hold date nation confirmed deaths recovered
. reshape huge confirmed deaths recovered, i(date) j(nation)
(word: j = 1 2 3)
Knowledge lengthy -> huge
------------------------------------------------------------------------
Variety of obs. 177 -> 62
Variety of variables 5 -> 10
j variable (3 values) nation -> (dropped)
xij variables:
confirmed -> confirmed1 confirmed2 confirmed3
deaths -> deaths1 deaths2 deaths3
recovered -> recovered1 recovered2 recovered3
------------------------------------------------------------------------
The output tells us that reshape modified the variety of observations from 177 in our unique dataset to 62 in our new dataset. We had 5 variables in our unique dataset, and we now have 10 variables in our new dataset. The variable nation in our outdated dataset has been faraway from our new dataset. The variable confirmed in our unique dataset has been mapped to the variables confirmed1, confirmed2, and confirmed3 in our new dataset. The variables deaths and recovered have been handled the identical manner. Let’s describe our new, huge dataset.
. describe
Comprises knowledge
obs: 62
vars: 10
------------------------------------------------------------------------
storage show worth
variable title sort format label variable label
------------------------------------------------------------------------
date float %td..
confirmed1 double %8.0g 1 confirmed
deaths1 double %8.0g 1 deaths
recovered1 double %8.0g 1 recovered
confirmed2 double %8.0g 2 confirmed
deaths2 double %8.0g 2 deaths
recovered2 double %8.0g 2 recovered
confirmed3 double %8.0g 3 confirmed
deaths3 double %8.0g 3 deaths
recovered3 double %8.0g 3 recovered
------------------------------------------------------------------------
Sorted by: date
The information for confirmed circumstances in China, Italy, and the USA in our unique dataset have been positioned, respectively, in variables confirmed1, confirmed2, and confirmed3 in our new dataset. How do I do know this?
Recall that the info for nation have been saved in a labeled, numeric variable. China was saved as 1, Italy was saved as 2, and the USA was saved as 3.
. label checklist nation
nation:
1 China
2 Italy
3 US
The numbers inform us which new variable goes with which nation. Let’s checklist the huge knowledge to confirm this sample.
. checklist date confirmed1 confirmed2 confirmed3 ///
in -5/l, abbreviate(13)
+---------------------------------------------------+
| date confirmed1 confirmed2 confirmed3 |
|---------------------------------------------------|
58. | 03/19/2020 81156 41035 13680 |
59. | 03/20/2020 81250 47021 19101 |
60. | 03/21/2020 81305 53578 25493 |
61. | 03/22/2020 81397 59138 33276 |
62. | 03/23/2020 81496 63927 43667 |
+---------------------------------------------------+
These variable names may very well be complicated, so let’s rename and label our variables to keep away from confusion. I’ll use the suffix _c to point “confirmed circumstances”, _d to point “deaths”, and _r to “point out recovered”. The variable labels will make this naming conference specific.
rename confirmed1 china_c
rename deaths1 china_d
rename recovered1 china_r
label var china_c "China circumstances"
label var china_d "China deaths"
label var china_r "China recovered"
rename confirmed2 italy_c
rename deaths2 italy_d
rename recovered2 italy_r
label var italy_c "Italy circumstances"
label var italy_d "Italy deaths"
label var italy_r "Italy recovered"
rename confirmed3 usa_c
rename deaths3 usa_d
rename recovered3 usa_r
label var usa_c "USA circumstances"
label var usa_d "USA deaths"
label var usa_r "USA recovered"
Let’s describe and checklist our knowledge to confirm our outcomes.
. describe
Comprises knowledge
obs: 62
vars: 10
------------------------------------------------------------------------
storage show worth
variable title sort format label variable label
------------------------------------------------------------------------
date float %td..
china_c double %8.0g China circumstances
china_d double %8.0g China deaths
china_r double %8.0g China recovered
italy_c double %8.0g Italy circumstances
italy_d double %8.0g Italy deaths
italy_r double %8.0g Italy recovered
usa_c double %8.0g USA circumstances
usa_d double %8.0g USA deaths
usa_r double %8.0g USA recovered
------------------------------------------------------------------------
Sorted by: date
. checklist date china_c italy_c usa_c ///
in -5/l, abbreviate(13)
+----------------------------------------+
| date china_c italy_c usa_c |
|----------------------------------------|
58. | 03/19/2020 81156 41035 13680 |
59. | 03/20/2020 81250 47021 19101 |
60. | 03/21/2020 81305 53578 25493 |
61. | 03/22/2020 81397 59138 33276 |
62. | 03/23/2020 81496 63927 43667 |
+----------------------------------------+
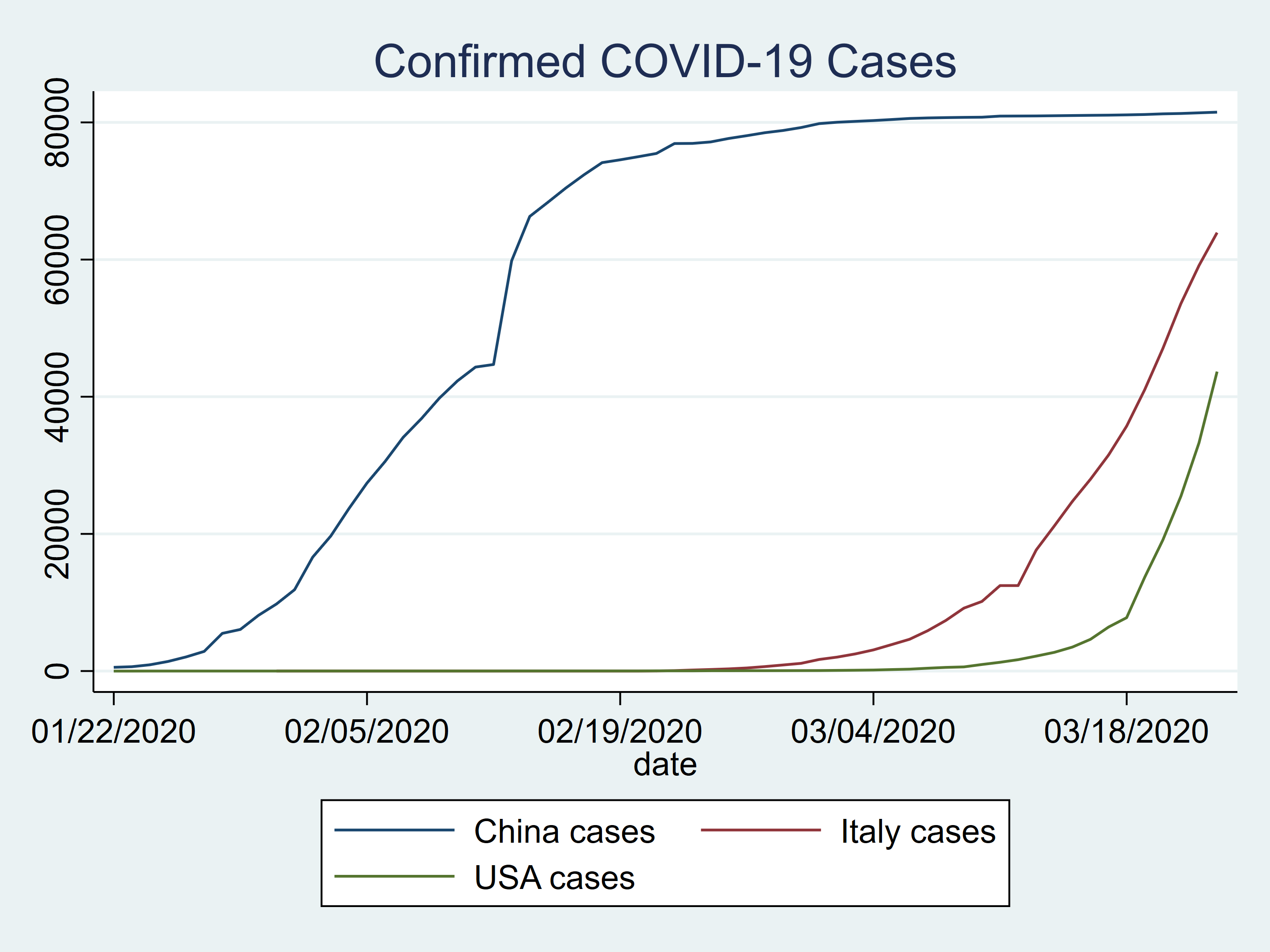
We may plot our knowledge to check the variety of confirmed circumstances for China, Italy, and the US.
twoway (line china_c date) ///
(line italy_c date) ///
(line usa_c date) ///
, title(Confirmed COVID-19 Circumstances)
Determine 2: Confirmed COVID-19 circumstances in China, Italy, and the US
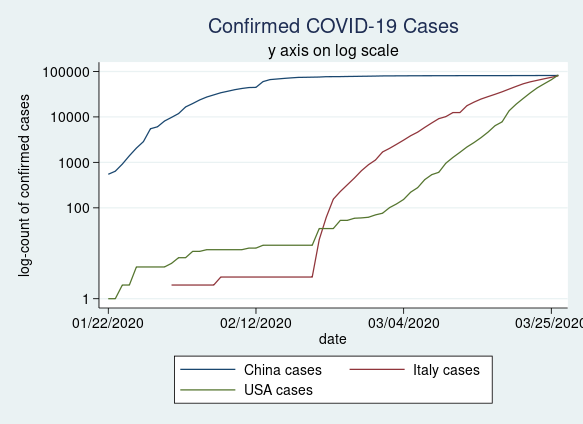
Many individuals choose to plot their knowledge on a log scale. That is straightforward to do utilizing the yscale(log) choice in our twoway command.
twoway (line china_c date) ///
(line italy_c date) ///
(line usa_c date) ///
, title(Confirmed COVID-19 Circumstances) ///
subtitle(y axis on log scale) ///
ytitle(log-count of confirmed circumstances) ///
yscale(log) ///
ylabel(0 100 1000 10000 100000, angle(horizontal))
Determine 3: Confirmed COVID-19 circumstances in China, Italy, and the USA plotted on a log scale
Utilizing uncooked counts could be deceptive as a result of the populations of China, Italy, and the USA are fairly totally different. Let’s create new variables that comprise the variety of confirmed circumstances per million inhabitants. The inhabitants knowledge come from the United States Census Burea Inhabitants Clock.
generate china_ca = china_c / 1389.6
generate italy_ca = italy_c / 62.3
generate usa_ca = usa_c / 331.8
label var china_ca "China circumstances adj."
label var italy_ca "Italy circumstances adj."
label var usa_ca "USA circumstances adj."
format %9.0f china_ca italy_ca usa_ca
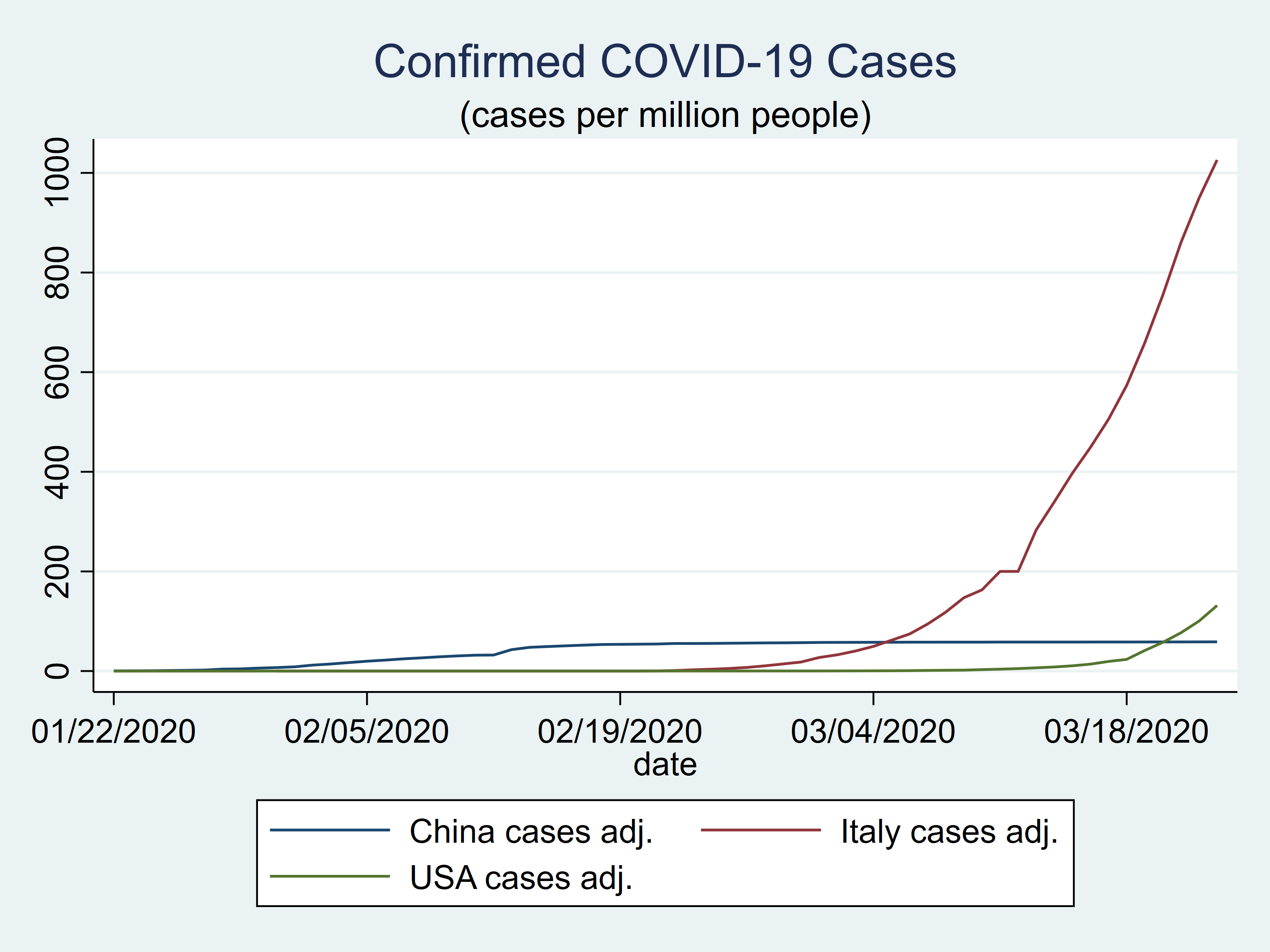
The plot of the population-adjusted knowledge seems fairly totally different from the plot of the unadjusted knowledge.
twoway (tsline china_ca) ///
(tsline italy_ca) ///
(tsline usa_ca) ///
, title(Confirmed COVID-19 Circumstances) ///
subtitle("(circumstances per million folks)")
Determine 4: Confirmed COVID-19 circumstances in China, Italy, and the USA adjusted for inhabitants dimension
We will add notes to our dataset to doc the calculations for the population-adjusted knowledge and the supply of the inhabitants knowledge.
notes china_ca: china_ca = china_c / 1389.6
notes china_ca: Inhabitants knowledge supply: https://www.census.gov/popclock/
notes italy_ca: italy_ca = italy_c / 62.3
notes italy_ca: Inhabitants knowledge supply: https://www.census.gov/popclock/
notes usa_ca: usa_ca = usa_c / 331.8
notes usa_ca: Inhabitants knowledge supply: https://www.census.gov/popclock/
We will view the notes by typing notes.
. notes china_ca: 1. china_ca = china_c / 1389.6 2. Inhabitants knowledge supply: https://www.census.gov/popclock/ italy_ca: 1. italy_ca = italy_c / 62.3 2. Inhabitants knowledge supply: https://www.census.gov/popclock/ usa_ca: 1. usa_ca = usa_c / 331.8 2. Inhabitants knowledge supply: https://www.census.gov/popclock/
We will additionally label our dataset and add notes for the complete dataset.
label knowledge "COVID-19 Knowledge assembled for the Stata Weblog"
notes _dta: Uncooked knowledge course: https://github.com/CSSEGISandData/COVID-19/tree/grasp/csse_covid_19_data/csse_covid_19_daily_reports
notes _dta: These knowledge are for educational functions solely
Final, we are able to tsset and save our dataset.
. tsset date, day by day
time variable: date, 01/22/2020 to 03/23/2020
delta: 1 day
. save covid19_wide, change
file covid19_wide.dta saved
Conclusion and mixed instructions
We did it! We efficiently downloaded the uncooked knowledge information, merged them, formatted them, and created two datasets that we are able to use to make tables and graphs. It will have been simpler if the info have been formatted persistently over time, however that’s the nature of actual knowledge. Happily, we now have the instruments and abilities we have to deal with these sorts of duties. I’ve collected the Stata instructions beneath so you’ll be able to run them should you like. The uncooked knowledge could change once more sooner or later, and it’s possible you’ll want to switch the code beneath to deal with these adjustments.
I might once more prefer to strongly emphasize that we now have not checked and cleaned these knowledge. The code and the ensuing knowledge needs to be used for educational functions solely.
Stata code to obtain COVID-19 knowledge from Johns Hopkins College as of March 23, 2020
native URL = "https://uncooked.githubusercontent.com/CSSEGISandData/COVID-19/grasp/csse_covid_19_data/csse_covid_19_daily_reports/"
forvalues month = 1/12 {
forvalues day = 1/31 {
native month = string(`month', "%02.0f")
native day = string(`day', "%02.0f")
native 12 months = "2020"
native right this moment = "`month'-`day'-`12 months'"
native FileName = "`URL'`right this moment'.csv"
clear
seize import delimited "`FileName'"
seize affirm variable ïprovincestate
if _rc == 0 {
rename ïprovincestate provincestate
label variable provincestate "Province/State"
}
seize rename province_state provincestate
seize rename country_region countryregion
seize rename last_update lastupdate
seize rename lat latitude
seize rename lengthy longitude
generate tempdate = "`right this moment'"
seize save "`right this moment'", change
}
}
clear
forvalues month = 1/12 {
forvalues day = 1/31 {
native month = string(`month', "%02.0f")
native day = string(`day', "%02.0f")
native 12 months = "2020"
native right this moment = "`month'-`day'-`12 months'"
seize append utilizing "`right this moment'"
}
}
generate date = date(tempdate, "MDY")
format date %tdNN/DD/CCYY
change countryregion = "China" if countryregion=="Mainland China"
hold if inlist(countryregion, "China", "US", "Italy")
collapse (sum) confirmed deaths recovered, by(date countryregion)
encode countryregion, gen(nation)
tsset nation date, day by day
save covid19_long, change
hold date nation confirmed deaths recovered
reshape huge confirmed deaths recovered, i(date) j(nation)
rename confirmed1 china_c
rename deaths1 china_d
rename recovered1 china_r
label var china_c "China circumstances"
label var china_d "China deaths"
label var china_r "China recovered"
rename confirmed2 italy_c
rename deaths2 italy_d
rename recovered2 italy_r
label var italy_c "Italy circumstances"
label var italy_d "Italy deaths"
label var italy_r "Italy recovered"
rename confirmed3 usa_c
rename deaths3 usa_d
rename recovered3 usa_r
label var usa_c "USA circumstances"
label var usa_d "USA deaths"
label var usa_r "USA recovered"
generate china_ca = china_c / 1389.6
generate italy_ca = italy_c / 62.3
generate usa_ca = usa_c / 331.8
label var china_ca "China circumstances adj."
label var italy_ca "Italy circumstances adj."
label var usa_ca "USA circumstances adj."
format %9.0f china_ca italy_ca usa_ca
notes china_ca: china_ca = china_c / 1389.6
notes china_ca: Inhabitants knowledge supply: https://www.census.gov/popclock/
notes italy_ca: italy_ca = italy_c / 62.3
notes italy_ca: Inhabitants knowledge supply: https://www.census.gov/popclock/
notes usa_ca: usa_ca = usa_c / 331.8
notes usa_ca: Inhabitants knowledge supply: https://www.census.gov/popclock/
label knowledge "COVID-19 Knowledge assembled for the Stata Weblog"
notes _dta: Uncooked knowledge course: https://github.com/CSSEGISandData/COVID-19/tree/grasp/csse_covid_19_data/csse_covid_19_daily_reports
notes _dta: These knowledge are for educational functions solely
tsset date, day by day
save covid19_wide, change